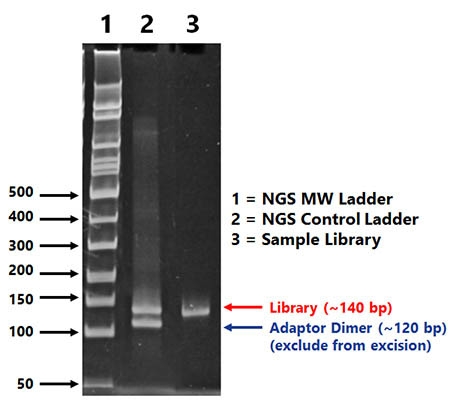
Randomisation of adapter sequences close to the ligation junction would neutralize this effect and improve the fidelity of NGS results. We will refer to this type of bias as “sequence bias”. Several studies demonstrated that adapter ligation steps may cause serious bias, due to RNA sequence/structure effects resulting in the preferential ligation of certain sRNAs with a given adapter sequence, while others are disfavoured. This modification strongly reduces the efficiency of 3′ adapter ligation, thus making library preparation particularly challenging for these types of sRNA. An additional problem arises with plant miRNAs, piRNAs in insects nematodes and mammals, and siRNAs in insects and plants in which the 3′ terminal nucleotide carries a 2’-O-methyl (2’ OMe) modification. Strongly underrepresented sRNAs, especially when their actual expression levels are already low, may remain undetected. Consequently, read numbers may not reflect actual sRNA expression levels and different sRNAs may be either over- or underrepresented in the library. All these steps are potential sources of bias. In a typical sRNA library preparation process, adapters are ligated to the RNAs followed by reverse transcription and PCR amplification. However, NGS approaches also have some disadvantages, such as the cost per sequencing run and the extensive processing steps required to convert a sample into a library for sequencing. In addition, it allows for the detection of single base differences and has a higher throughput than qPCR or Northern blotting. It allows for the discovery of novel sRNAs, has a better signal to noise ratio than microarrays and does not suffer of saturation effects. It has several advantages over microarray techniques or quantitative PCR (qPCR). Next-generation sequencing (NGS) is a powerful tool for the analysis of sRNAs. Altered miRNA expression profiles have been implicated in a number of diseases, highlighting the importance of miRNAs in biology and the need for continued development of research tools for the study of sRNA in general. Eukaryotic regulatory sRNAs typically range in size from ~ 20 to 30 nt and the three major classes are microRNAs (miRNA), small interfering RNAs (siRNA) and piwi-interacting RNAs (piRNA). Small RNAs (sRNAs) are known to play an important regulatory role in a wide range of organisms in many biological processes including embryo development, cell differentiation, growth/proliferation and apoptosis/cell death. In addition, we modified the TruSeq and NEXTflex protocols to identify conditions that improve performance. Here we compared these methods with Illumina’s classical TruSeq protocol regarding the detection of normal and 2’ OMe RNAs. To reduce bias, the NEBNext kit (New England Biolabs) uses polyethylene glycol (PEG), the NEXTflex V2 kit (BIOO Scientific) uses both randomised adapters and PEG, and the novel SMARTer (Clontech) and CATS (Diagenode) kits avoid ligation altogether. This inhibits 3′ adapter ligation and makes library preparation particularly challenging. Several types of sRNA including plant microRNAs (miRNA), piwi-interacting RNAs (piRNA) in insects, nematodes and mammals, and small interfering RNAs (siRNA) in insects and plants contain a 2’-O-methyl (2’-OMe) modification at their 3′ terminal nucleotide. However, classical sRNA library preparation methods introduce serious bias, mainly during adapter ligation steps. Next-generation sequencing technologies have revolutionized the study of small RNAs (sRNAs) on a genome-wide scale. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
